Author: Denis Avetisyan
Researchers are exploring how deep neural networks can learn to perform statistical inference directly from simulated data, bypassing the need for complex likelihood calculations.

This work introduces ForwardFlow, a simulation-based inference method leveraging deep normalizing flows to approximate sufficient statistics for parametric models, offering advantages in robustness and speed.
Traditional statistical inference often relies on analytically tractable likelihoods, a limitation increasingly challenged by complex models. The work presented in ‘ForwardFlow: Simulation only statistical inference using deep learning’ introduces a novel simulation-based approach leveraging deep neural networks to directly learn sufficient statistics for parameter estimation. By training on simulated data, this ‘forward’ method bypasses explicit likelihood specification, offering robustness to data contamination and algorithmic approximation errors. Could this paradigm shift unlock efficient inference in scenarios where traditional methods falter, and pave the way for pre-trained models applicable across diverse scientific domains?
The Fragility of Traditional Statistical Foundations
Traditional statistical inference frequently hinges on assumptions regarding the underlying distribution of data – often normality or other well-defined forms. However, real-world phenomena rarely conform to these idealized conditions. Complex systems exhibit non-linear relationships, skewed distributions, and dependencies that invalidate the core tenets of many conventional methods. For instance, modeling financial markets or ecological populations requires accounting for factors like volatility clustering or species interactions, which routinely violate assumptions of independence and constant variance. Consequently, applying standard techniques without careful consideration of these deviations can lead to inaccurate estimations, misleading conclusions, and an underestimation of inherent uncertainty, highlighting the need for more flexible and robust analytical approaches.
Frequentist statistical inference, a cornerstone of many scientific disciplines, encounters significant challenges when faced with datasets of limited size. This isn’t simply a matter of reduced statistical power; the very foundations of these methods – relying on the long-run frequency of events – become shaky with insufficient observations. Confidence intervals, for example, may fail to accurately reflect the true uncertainty surrounding an estimate, often appearing narrower and more precise than justified. Consequently, researchers may draw definitive conclusions from data that doesn’t support them, leading to inflated Type I error rates – incorrectly rejecting a true null hypothesis. The problem is compounded by the difficulty in accurately estimating parameters necessary for the correct application of these techniques; small errors in these estimates can dramatically skew results, particularly in complex models. Ultimately, the reliability of frequentist conclusions hinges on the availability of ample data, and its limitations must be carefully considered when interpreting findings from studies with modest sample sizes.
While Bayesian modeling presents a compelling alternative to traditional statistical inference, its practical application is often hampered by substantial computational demands. The core of Bayesian analysis involves calculating the posterior probability-the updated belief about a hypothesis given observed data-which requires integrating over all possible parameter values. This integration becomes exceedingly difficult, if not impossible, to perform analytically for models with numerous parameters or complex relationships. Consequently, researchers frequently rely on computationally intensive methods like Markov Chain Monte Carlo (MCMC) simulations to approximate the posterior distribution. These simulations, while effective, can require significant processing time and resources, particularly when dealing with high-dimensional data or intricate model structures, limiting the scalability and accessibility of Bayesian approaches for certain real-world problems. The pursuit of efficient Bayesian computation remains a critical area of research, driving the development of novel algorithms and approximation techniques to overcome these limitations.
A Shift in Perspective: Simulation-Only Inference
Simulation-Only Inference (SOI) represents a departure from traditional statistical methods by eliminating the requirement for an explicitly defined likelihood function. This is achieved through the generation of synthetic data from a model, allowing inference to be performed by comparing observed data to this simulated distribution. The avoidance of likelihood specification enhances robustness to model misspecification, as the method relies less on the accuracy of the assumed data-generating process. Consequently, SOI is particularly advantageous when dealing with complex models where defining or accurately estimating a likelihood function is computationally challenging or analytically intractable, offering increased flexibility in scenarios involving non-standard data distributions or model structures.
Simulation-only inference streamlines the statistical estimation process by employing a single neural network to learn sufficient statistics directly from the observed data. This contrasts with traditional methods requiring the specification of explicit likelihood functions and often extensive parameter tuning. The network effectively compresses the raw data into a lower-dimensional representation that captures all relevant information for inference, thereby reducing the computational demands associated with both model training and subsequent simulation runs. Consequently, the need for manual hyperparameter optimization is significantly lessened, as the network learns to extract the necessary features for accurate estimation, leading to a more automated and efficient inference pipeline.
Rao-Blackwellization is a technique used within simulation-only inference to enhance the precision of estimators derived from simulated data. This method involves conditioning the estimator on a sufficient statistic of the observed data, effectively reducing its variance without introducing bias. By leveraging the properties of sufficient statistics, Rao-Blackwellization constructs a new estimator that is, on average, closer to the true parameter value than the original estimator. This is achieved by replacing the original estimator with its conditional expectation given the sufficient statistic, thereby minimizing the mean squared error and improving the efficiency of the simulation process. The technique is particularly valuable when dealing with complex models where direct calculation of the likelihood function is intractable.
Architectural Foundations for Efficient Simulation
Branched network structures, in the context of simulation, facilitate parallel exploration of the parameter space by dividing the overall simulation task into multiple independent branches. Each branch operates on a subset of the parameter space, allowing for concurrent computation and a reduction in total simulation time. This approach contrasts with sequential methods where parameter space exploration occurs linearly. The degree of branching-the number of concurrent branches-can be adjusted to balance computational resources and the desired level of parallelism. While increasing the number of branches generally accelerates the process, it also introduces overhead associated with managing and coordinating these parallel computations, necessitating a careful optimization of the branching factor for maximal efficiency.
Coordinate-wise dense layers, applied within branched network structures, perform feature extraction on each dimension of the input parameter space independently, increasing representational capacity. These layers are followed by collapsing layers which reduce dimensionality by combining features across these independent dimensions. This process utilizes linear transformations and potentially non-linear activation functions to project high-dimensional input data into a lower-dimensional space while preserving relevant information for subsequent simulation steps. The combined effect is a more efficient parameterization and a reduction in computational cost associated with exploring the simulation space, as the number of parameters requiring optimization is decreased.
Deep Neural Networks (DNNs) facilitate scalable inference by learning sufficient statistics from high-dimensional parameter spaces. Traditional methods for parameter estimation often rely on exhaustive search or simplified models, becoming computationally intractable as dimensionality increases. DNNs, however, can approximate complex, non-linear functions, enabling them to learn a compressed representation of the parameter space – these learned statistics capture the essential information needed for inference. This allows for the estimation of parameter distributions and the prediction of simulation outcomes without explicitly calculating every possible parameter combination. The network’s ability to generalize from a limited set of training data to unseen parameter values is crucial for achieving scalability in simulations, particularly those involving a large number of degrees of freedom. \hat{\theta} = DNN(X) represents the approximated parameter estimator \hat{\theta} derived from input data X by a Deep Neural Network.
Safeguarding Robustness in the Face of Imperfection
Simulation-only inference offers a powerful pathway to robust statistical conclusions, particularly when facing the increasing threat of data contamination-outliers, errors, or even malicious manipulation within datasets. Unlike traditional methods heavily reliant on distributional assumptions, these approaches assess the plausibility of models by directly comparing observed data to a vast landscape of simulated outcomes. This inherently provides protection; a single, or even a few, corrupted data points exert minimal influence on the overall simulation-based assessment. The focus shifts from precise parameter estimation to determining whether the observed data could reasonably have arisen from the proposed model, effectively sidestepping the vulnerabilities that plague methods susceptible to undue influence from compromised observations. Consequently, simulation-only inference provides a reliable framework for drawing valid conclusions even in the presence of imperfect or adversarial data.
Integrating the Expectation-Maximization (EM) algorithm within a simulation-based inference pipeline offers a powerful strategy for refining parameter estimates. This iterative process begins by using the simulation to generate data based on initial parameter guesses, then leverages the observed data to update those guesses through expectation and maximization steps. The expectation step calculates the likely values of latent variables, while the maximization step adjusts the parameters to best fit the observed data given those estimated latent variables. This cyclical process continues until convergence, resulting in more precise and robust parameter estimates than those achievable through simulation alone. By combining the exploratory power of simulation with the refinement capabilities of the EM algorithm, researchers can effectively navigate complex models and extract meaningful insights from their data, even in the presence of uncertainty or missing information.
Computational methods for robust inference increasingly rely on techniques like Importance Sampling and Approximate Bayesian Computation (ABC) to overcome the limitations of traditional statistical approaches. These methods substantially improve the efficiency of simulations by focusing computational effort on the most informative regions of the parameter space. Recent applications to genetic data demonstrate the power of this combination, achieving a high coverage probability of 0.942 – indicating that the true parameter values are accurately captured within the estimated credible intervals – alongside a remarkably low root mean squared error (rMSE) of 0.01. This level of precision suggests these techniques offer a reliable pathway to robust and accurate parameter estimation even in complex datasets where conventional methods may struggle.
Charting a Course for Future Inference
Normalizing flows represent a powerful departure from traditional Approximate Bayesian Computation (ABC) methods by employing deep neural networks to learn a complex, invertible mapping between a simple probability distribution and the target posterior distribution. This approach fundamentally alters how inference is performed; rather than relying on summary statistics and acceptance thresholds, normalizing flows directly model the posterior, enabling efficient sampling and density estimation. By learning this normalizing mapping, these techniques sidestep the need for computationally expensive likelihood evaluations, accelerating the inference process and allowing researchers to tackle more intricate statistical problems. The method’s efficiency stems from the network’s ability to transform samples from a known distribution into samples that accurately represent the desired posterior, thereby circumventing the limitations of rejection-based ABC algorithms and offering a substantial gain in computational speed.
The convergence of normalizing flows and simulation-only inference represents a substantial advancement in addressing intricate statistical challenges. By bypassing the need for explicit likelihood functions – often intractable for complex models – researchers can now generate datasets directly from a prior, leveraging the power of deep learning to map these simulations to observed data. This approach not only streamlines the inference process but also unlocks the potential to explore high-dimensional parameter spaces previously inaccessible due to computational limitations. The resulting flexibility allows for the development of more nuanced and realistic models, capable of capturing subtle relationships within data and offering improved predictive power across a range of scientific disciplines, from genetics and epidemiology to cosmology and materials science.
A compelling advantage of this methodological shift lies in its drastically reduced implementation complexity. Current genotype simulation techniques, often reliant on Expectation-Maximization (EM) algorithms, typically demand approximately 100 lines of code to achieve comparable results. In contrast, the proposed normalizing flow approach accomplishes the same task with a remarkably concise 10 lines of code. This simplification not only accelerates the development process but also enhances code readability and maintainability, lowering the barrier to entry for researchers and facilitating broader adoption of sophisticated statistical inference methods in genetics and related fields. The efficiency gain promises to accelerate research cycles and empower scientists to focus on biological insights rather than intricate coding challenges.
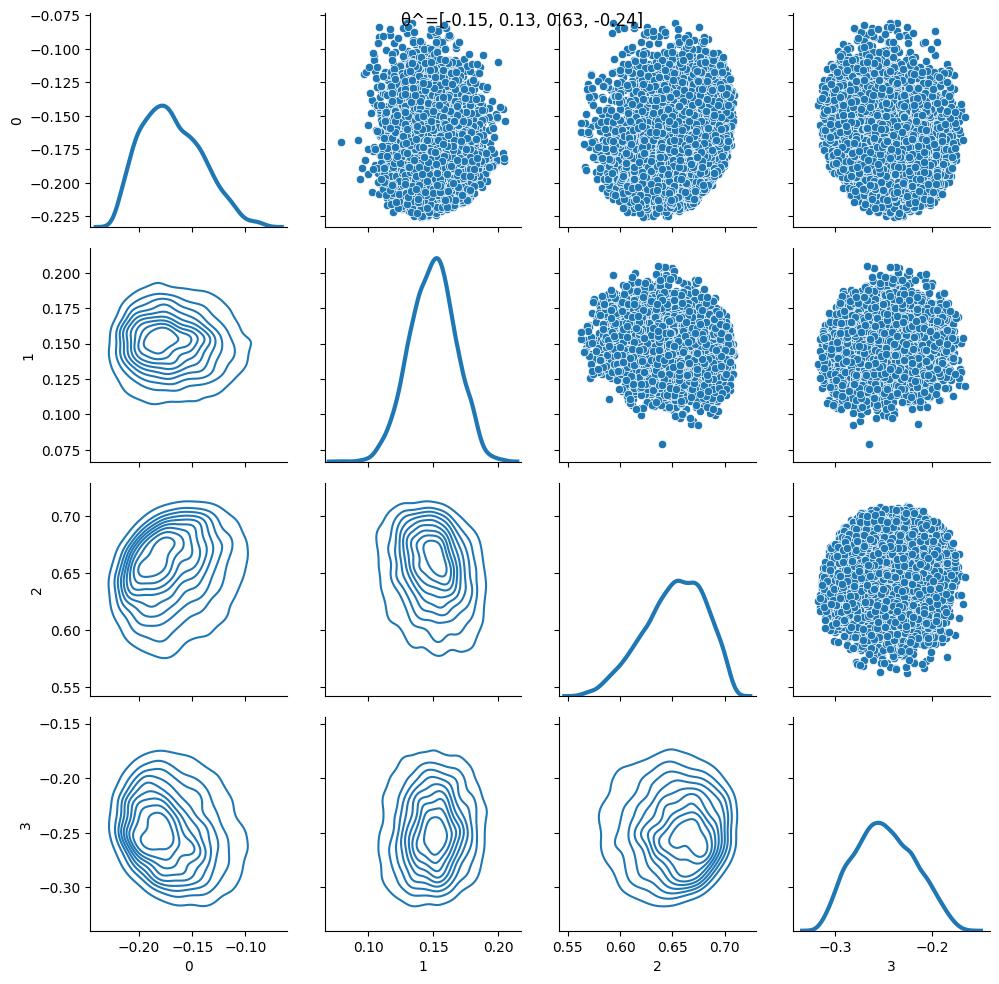
The pursuit of efficient statistical inference, as detailed in this work, echoes a fundamental principle of system design: structure dictates behavior. ForwardFlow’s approach, by learning sufficient statistics directly from simulation, bypasses the often-fragile assumptions inherent in traditional methods. This reliance on learned structure, rather than pre-defined likelihoods, offers resilience against data contamination – a critical consideration in modern data science. As Sergey Sobolev once stated, “The essence of a good model is not to predict the future, but to understand the present.” This sentiment perfectly captures the core of ForwardFlow; the system aims to accurately represent the underlying data-generating process, thereby enabling robust inference even in challenging scenarios. Good architecture is invisible until it breaks, and only then is the true cost of decisions visible.
Where Do the Currents Take Us?
The elegance of ForwardFlow lies in its deliberate refusal to explicitly model the likelihood – a function often approximated, massaged, and ultimately, strained by the realities of complex data. However, this avoidance begs a question: what are the true optimization targets when dispensing with likelihoods altogether? The paper rightly points toward sufficient statistics, but their identification remains, fundamentally, a modeling choice. Future work must rigorously examine the sensitivity of inference to misspecification of these statistics, and explore adaptive methods that learn them directly from the data – a truly end-to-end approach.
The resilience to data contamination is a compelling feature, yet it skirts a deeper issue. Any statistical method, simulation-based or otherwise, operates under assumptions – implicit or explicit – about the data generating process. A contaminated dataset isn’t simply ‘noisy’; it signals a model breakdown. The challenge isn’t merely to tolerate contamination, but to detect it, and to acknowledge the limitations of any analysis performed in its presence. Simplicity, in this context, isn’t about minimizing complexity, but about honestly acknowledging what the data cannot tell us.
Finally, the focus on parametric models, while practical, feels…limiting. The true power of simulation-based inference may lie in its potential to move beyond parameter estimation altogether – to explore model structures themselves, and to quantify uncertainty not just in parameter values, but in the very assumptions underlying the model. The currents are strong, but the destination remains uncharted.
Original article: https://arxiv.org/pdf/2603.10991.pdf
Contact the author: https://www.linkedin.com/in/avetisyan/
See also:
- Gold Rate Forecast
- The Best Switch RPGs to Play Using Switch 2 Handheld Boost Mode
- Avengers: Doomsday Spoilers & Leaks Addressed By Director Joe Russo: “It’s Over-Policed”
- INJ/USD
- STX/USD
- 5 Horror Shows I Knew Would Be 10/10 Masterpieces After The First 10 Minutes
- What is Omoggle? The AI face-rating platform taking over Twitch
- Lord Of The Flies Review: Near-Perfect Adaptation Is A Reminder Of Classic Novel’s Haunting Power
- Man pulls car with his manhood while on fire to raise awareness for prostate cancer
- Netflix’s Little House On The Prairie Reboot: Release Date, Cast & Everything We Know
2026-03-13 01:44