Supply Chain Domino Effects: Predicting Software Vulnerability Cascades
A new approach utilizes software bill of materials and advanced graph neural networks to anticipate how vulnerabilities can spread through complex software dependencies.
A new approach utilizes software bill of materials and advanced graph neural networks to anticipate how vulnerabilities can spread through complex software dependencies.
New research tackles the challenge of maintaining accurate credit risk assessments as customer behavior and economic conditions evolve.
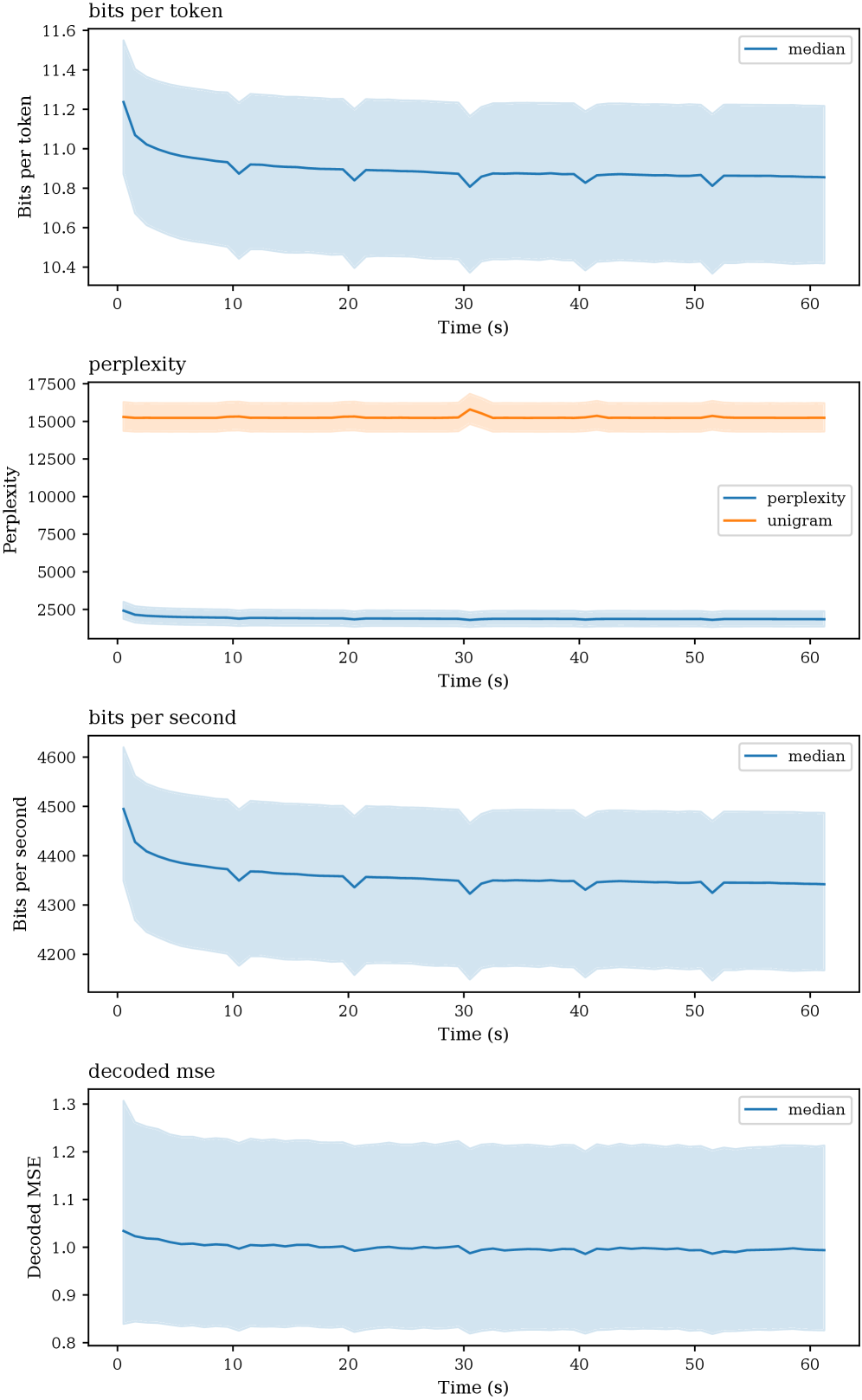
Researchers have developed a generative model capable of producing remarkably realistic magnetoencephalography (MEG) signals, opening new avenues for understanding and simulating brain activity.
![Assets in a two-dimensional principal component space reveal statistical outliers-specifically, BTC, GALA, and SC, exceeding a [latex]2\sigma[/latex] threshold-and demonstrate distinct clusters identified through cross-sectional analysis, suggesting inherent groupings within the asset landscape.](https://arxiv.org/html/2601.20336v1/x2.png)
New research casts doubt on the ability of cryptocurrency project whitepapers to accurately predict actual market performance.
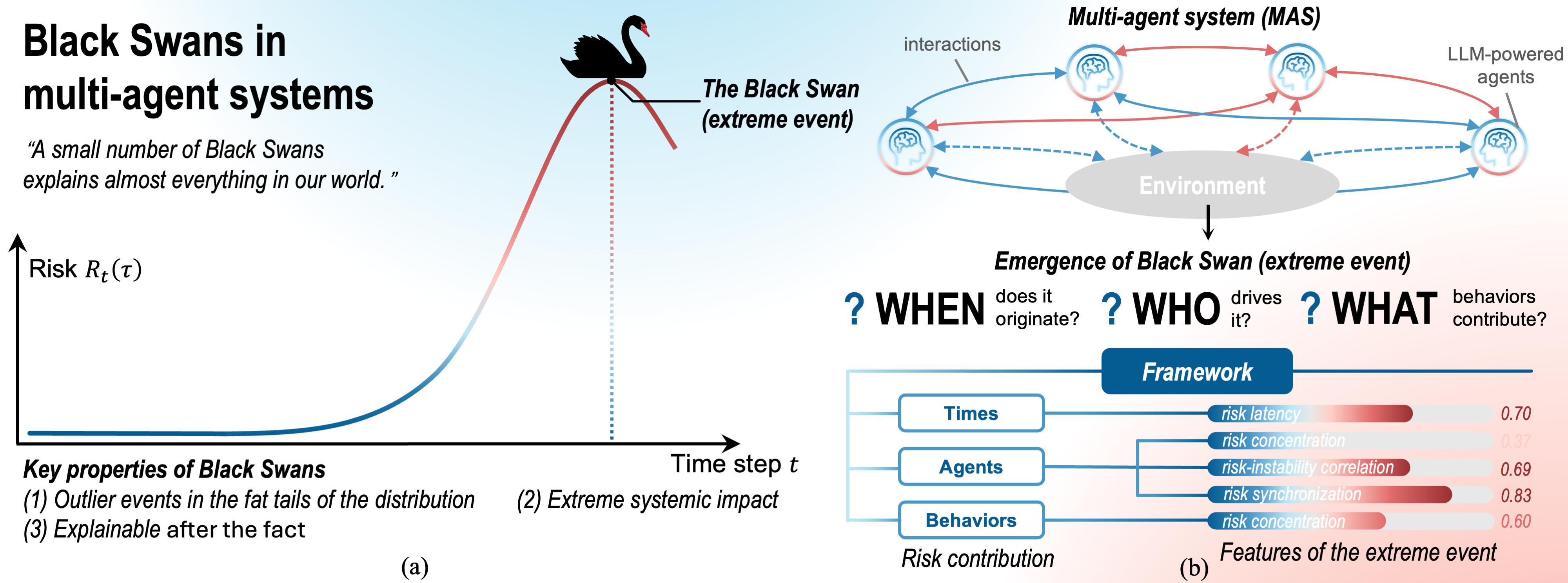
A new framework explains how unexpected failures arise in complex systems of AI agents, pinpointing the contributions of individual actors to catastrophic outcomes.
A new approach combines geodetic data with the physics of fault behavior, offering a powerful tool for understanding and potentially forecasting slow slip events.
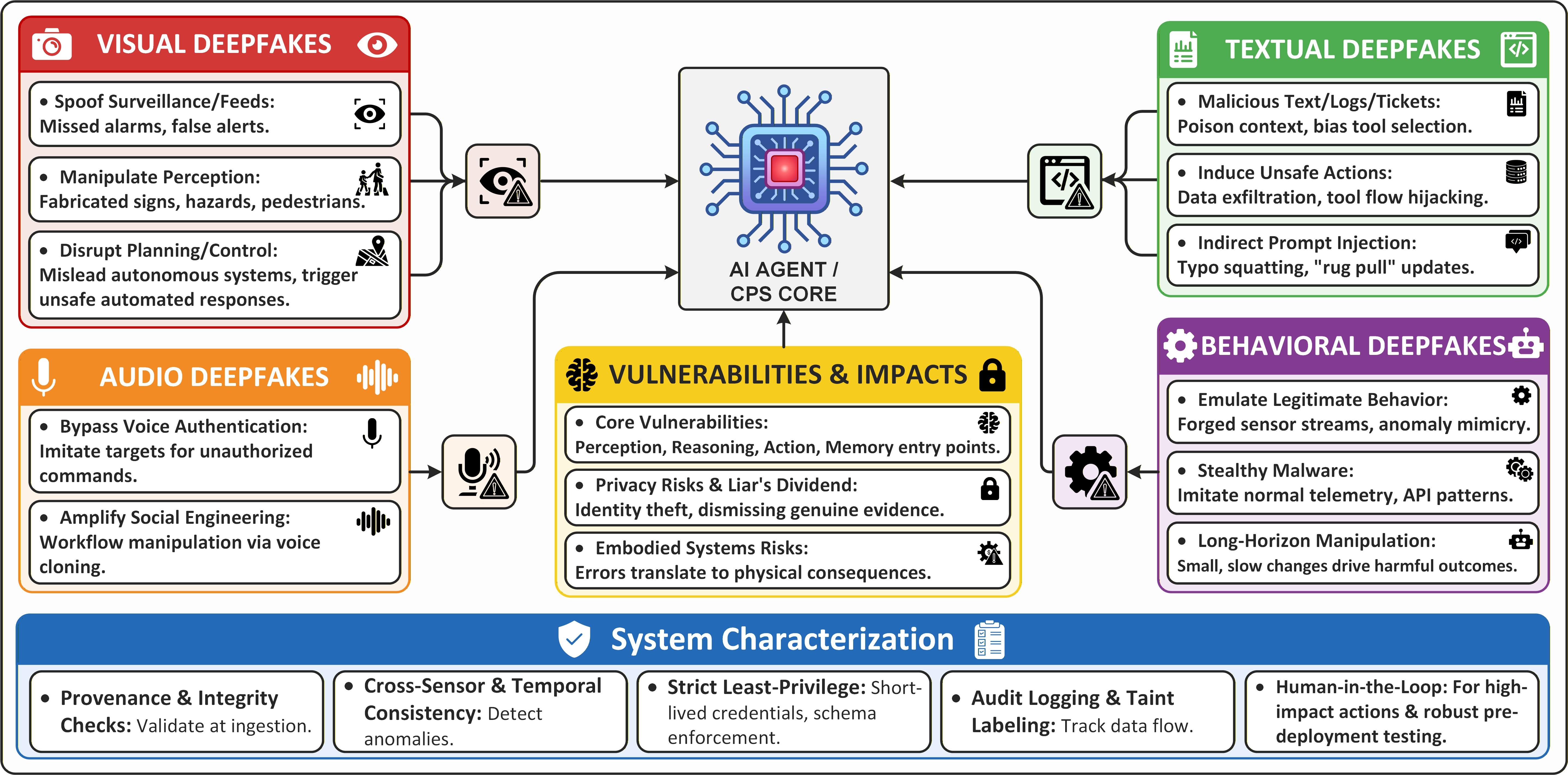
As artificial intelligence increasingly controls critical infrastructure, the potential for deception through manipulated sensory data presents a growing threat to the safety and reliability of cyber-physical systems.
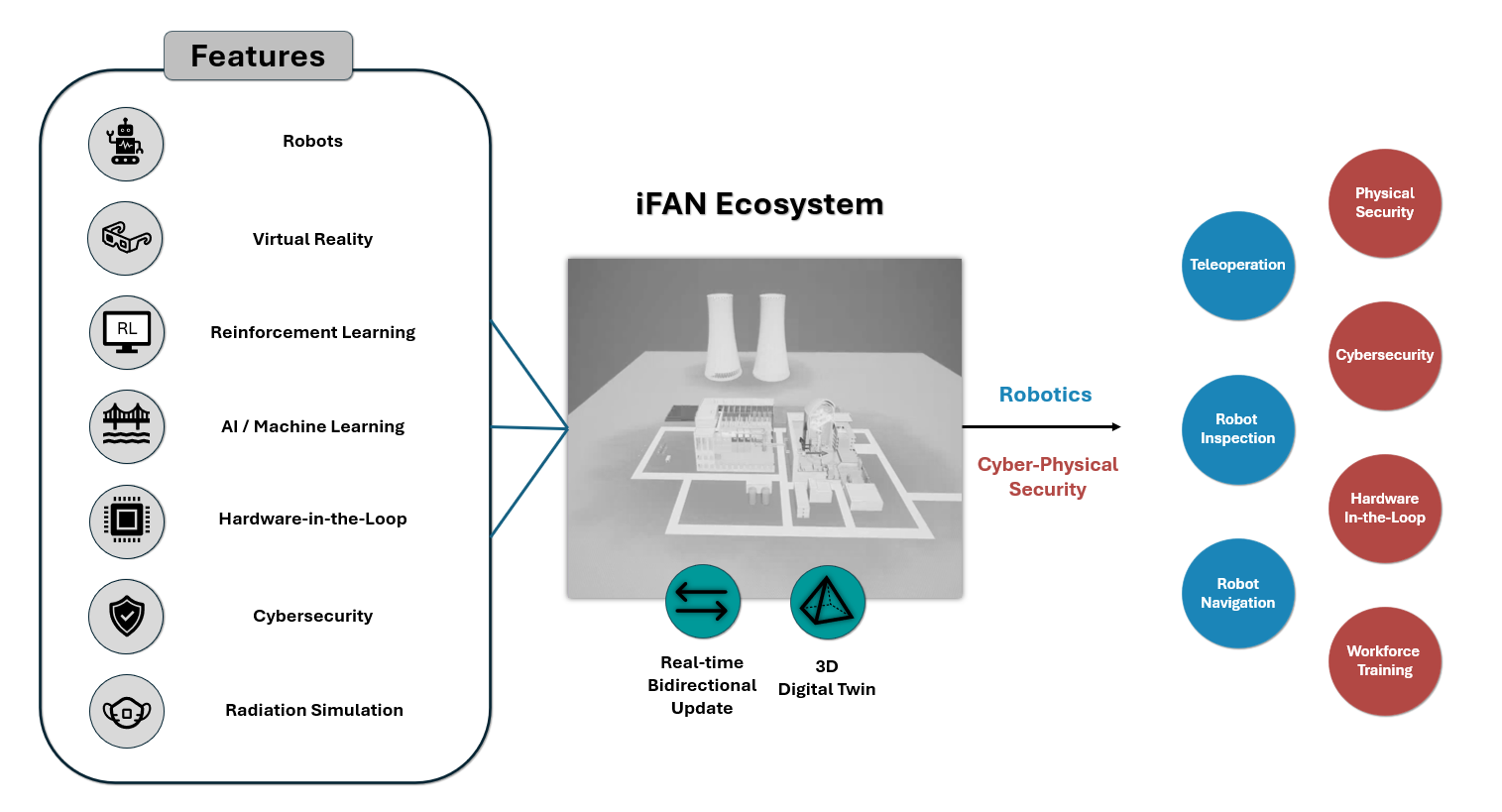
A new integrated digital twin environment is poised to accelerate innovation in nuclear plant technology and bolster operational safety through advanced robotics, AI, and cybersecurity.
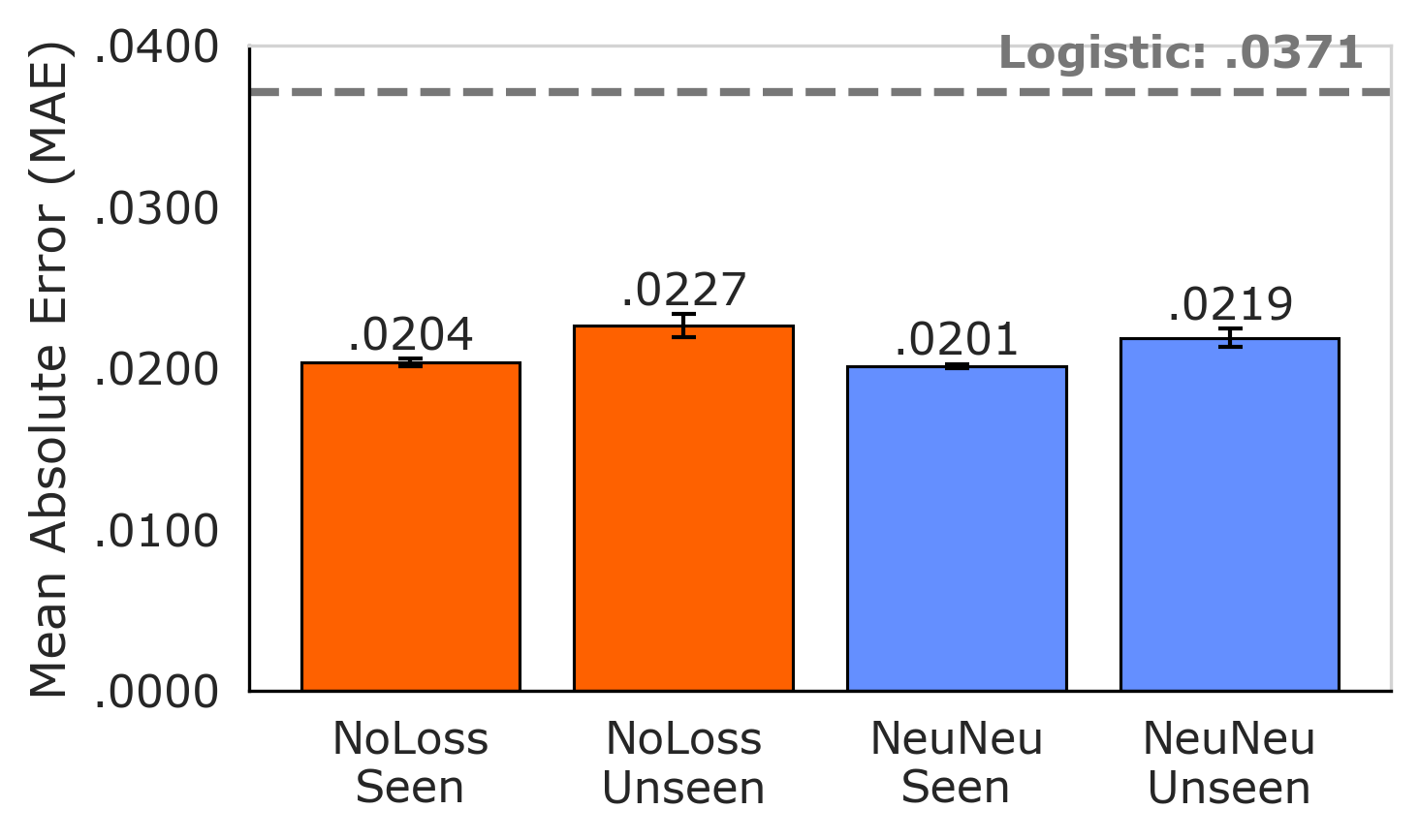
A new approach leverages token-level loss to more accurately forecast how well a model will perform on unseen tasks.
![The study demonstrates a transformer model’s capacity to accurately predict the short-term dynamics of a charge density wave (CDW) order parameter [latex]\Delta\_{\rho}(t)[/latex], effectively mirroring exact simulations, though inherent error accumulation within the chaotic regime leads to divergence over extended timescales-nevertheless, the model successfully captures the statistical characteristics of the system’s dynamic behavior.](https://arxiv.org/html/2601.19080v1/x4.png)
Researchers are leveraging the power of transformer networks to model and predict the collective behavior of chaotic many-body systems, opening new avenues for understanding complex phenomena.